Recent News
Economic Transformation and CIO Challenges: Middle Eastern Visionaries Driving Industry 4.0
Economic transformation in the Middle East is being driven by visionary Chief Information Officers (CIOs) who…
Economic Diversification and Arab CIOs: Transforming Industries with Emerging Technologies
Economic diversification is a strategic priority for many Arab countries as they seek to reduce reliance…
Navigating Digital Transformation: Middle Eastern CIOs’ Strategies and Roadblocks
Middle Eastern Chief Information Officers (CIOs) are at the forefront of navigating digital transformation in their…
E-Government Evolution: Arab CIOs Leading Digital Transformation in Public Services
Arab Chief Information Officers (CIOs) are at the forefront of the e-government evolution, leading the digital…
UAE TECH CEOS
ARTIFICIAL INTELLIGENCE
Exploring the Frontier: A Forward-Looking Analysis of Emerging AI Trends, Revolutionary Innovations, and Their Implications for Global Progress and Human Prosperity
As we venture into the future, the landscape of artificial intelligence (AI) continues to evolve at a rapid pace, ushering in a new era of…
Navigating the EMEA AI Ecosystem: Insights into Key Players, Collaborations, and Breakthroughs Fueling the Growth of Artificial Intelligence in Europe, the Middle East, and Africa
The EMEA region boasts a rich and diverse ecosystem driving the growth of artificial intelligence (AI) across Europe, the Middle East, and Africa. From established…
From Silicon Valleys to AI Oases: Spotlight on Emerging Tech Hubs in the EMEA Region Driving Innovation and Advancement in Artificial Intelligence
The EMEA region is witnessing a proliferation of emerging tech hubs that are becoming focal points for innovation and advancement in artificial intelligence (AI). These…
Unveiling the EMEA AI Revolution: Trends, Startups, and Investments Shaping the Future of Artificial Intelligence in Europe, the Middle East, and Africa
As the global AI landscape continues to evolve, the EMEA region stands out as a vibrant hub of innovation and investment in artificial intelligence. From…
AI Renaissance in the EMEA: Exploring Cutting-Edge Technologies and Innovations Transforming Industries Across Europe, the Middle East, and Africa
The AI renaissance in the EMEA region (Europe, the Middle East, and Africa) is marked by a surge in cutting-edge technologies and innovations that are…
The Next Chapter: Anticipating the Unfolding Narrative of AI Technology – Insights into Future Trends, Transformative Innovations, and the Journey Towards Artificial General Intelligence (AGI)
Abstract: Artificial Intelligence (AI) has entered a phase of unprecedented acceleration, reshaping industries, societies, and the very fabric of human existence. As we stand on…
Forging New Frontiers: A Holistic Perspective on the Evolving Landscape of Artificial Intelligence – Trends, Disruptions, and Opportunities in an Era of Rapid Technological Advancement
Abstract: In an era defined by rapid technological advancement, Artificial Intelligence (AI) stands as a pivotal force reshaping industries, economies, and societies at large. This…
Tomorrow’s Intelligence: A Deep Dive into the Future of AI Technology – From Advancements in Neural Networks to the Ethical Dimensions of Machine Learning Systems
Abstract: Artificial Intelligence (AI) technology continues to evolve at a rapid pace, shaping our future in profound ways. This paper provides an in-depth exploration of…
Embracing the Unknown: Exploring the Uncharted Territory of Tomorrow’s AI Technologies – A Multifaceted Examination of Trends, Innovations, and Implications
Abstract: The landscape of Artificial Intelligence (AI) is constantly evolving, presenting both unprecedented opportunities and daunting challenges. This paper embarks on a journey into the…
The Road Ahead: An In-Depth Analysis of Emerging AI Trends, from Autonomous Systems to Personalized Intelligence, and Their Role in Shaping Our Collective Future
Abstract: The evolution of Artificial Intelligence (AI) continues to accelerate, paving the way for transformative advancements in various domains. This paper provides a comprehensive analysis…
Beyond Imagination: Delving into the Future of AI Technologies and Their Potential to Reshape Humanity, Workforce Dynamics, and Societal Norms
Abstract: Artificial Intelligence (AI) technologies are poised to redefine the fabric of human society, fundamentally altering the way we work, interact, and perceive the world…
Towards a New Era: Anticipating the Future Trajectory of Artificial Intelligence – Trends, Challenges, and Promises on the Horizon
Abstract: Artificial Intelligence (AI) stands at the forefront of technological innovation, shaping industries, societies, and economies alike. As we stand on the cusp of a…
Frontiers of Possibility: Unveiling the Next Generation of AI Technology Trends, from Quantum Computing to Human-Machine Collaboration
Frontiers of Possibility: Unveiling the Next Generation of AI Technology Trends, from Quantum Computing to Human-Machine Collaboration" suggests an exploration of cutting-edge AI technology trends,…
Charting the Course: Navigating the Complex Landscape of Tomorrow’s AI Innovations, Trends, and Ethical Considerations
Navigating the complex landscape of tomorrow's AI innovations, trends, and ethical considerations requires a multi-faceted approach that integrates technological advancement with ethical foresight. Here's a…
Into the Unknown: A Comprehensive Exploration of Future AI Technologies and Their Implications for Industries, Economies, and Global Dynamics
Artificial Intelligence (AI) is poised to revolutionize industries, economies, and global dynamics in unprecedented ways. As AI technologies continue to advance at an exponential rate,…
Navigating the Shadows: Uncovering Ethical Concerns and Risks Associated with Artificial Intelligence
Artificial Intelligence (AI) has been rapidly advancing over the past few years, with numerous benefits and opportunities. However, as with any technology, there are also…
What We Offer:
Technlogy Stories
BIOTECH CEOS
BIOTECH
The European Bioeconomy: A Roadmap to Innovation, Competitiveness, and Sustainability – Policies, Investments, and Partnerships Driving Biotechnology Advancements and Economic Growth
"The European Bioeconomy: A Roadmap to Innovation, Competitiveness, and Sustainability - Policies, Investments, and Partnerships Driving Biotechnology Advancements and Economic Growth" suggests an exploration of…
Investing in Europe’s Biotech Future: Analyzing Funding Trends, Venture Capital Landscape, and Growth Opportunities in the Continent’s Dynamic Life Sciences Industry
"Investing in Europe's Biotech Future: Analyzing Funding Trends, Venture Capital Landscape, and Growth Opportunities in the Continent's Dynamic Life Sciences Industry" suggests an in-depth analysis…
Sustainable Biotech Solutions: Europe’s Contribution to Addressing Environmental Challenges, Sustainable Agriculture, and Circular Economy Principles Through Biotechnology Innovations
"Sustainable Biotech Solutions: Europe's Contribution to Addressing Environmental Challenges, Sustainable Agriculture, and Circular Economy Principles Through Biotechnology Innovations" suggests an exploration of Europe's efforts to…
Revolutionizing Healthcare: How Europe’s Biotech Sector is Redefining Diagnostics, Therapeutics, and Disease Prevention Through Innovations in Molecular Biology and Bioinformatics
"Revolutionizing Healthcare: How Europe's Biotech Sector is Redefining Diagnostics, Therapeutics, and Disease Prevention Through Innovations in Molecular Biology and Bioinformatics" suggests a comprehensive exploration of…
Biosecurity in Europe: Safeguarding Public Health, Biodiversity, and Food Systems – Strategies, Technologies, and Policy Imperatives in the Face of Emerging Biological Threats
"Biosecurity in Europe: Safeguarding Public Health, Biodiversity, and Food Systems - Strategies, Technologies, and Policy Imperatives in the Face of Emerging Biological Threats" suggests a…
Biotech Hubs of Europe: Spotlight on Clusters, Startups, and Research Institutions Driving Innovation and Collaboration in the Continent’s Thriving Life Sciences Ecosystem
"Biotech Hubs of Europe: Spotlight on Clusters, Startups, and Research Institutions Driving Innovation and Collaboration in the Continent's Thriving Life Sciences Ecosystem" suggests an in-depth…
Biotech Resilience in Europe: Navigating Regulatory Frameworks, Investment Trends, and Global Competitiveness in an Era of Scientific Advancement and Economic Uncertainty
"Biotech Resilience in Europe: Navigating Regulatory Frameworks, Investment Trends, and Global Competitiveness in an Era of Scientific Advancement and Economic Uncertainty" suggests an in-depth examination…
Genetic Frontiers: Europe’s Journey in Biotechnology – From CRISPR to Gene Therapy, A Comprehensive Analysis of Innovations, Regulations, and Ethical Considerations
"Genetic Frontiers: Europe's Journey in Biotechnology - From CRISPR to Gene Therapy, A Comprehensive Analysis of Innovations, Regulations, and Ethical Considerations" suggests a detailed exploration…
Europe’s Biotech Renaissance: Unveiling the Continent’s Role in Advancing Cutting-Edge Therapies, Precision Medicine, and Sustainable Solutions for Global Health Challenges
"Europe's Biotech Renaissance: Unveiling the Continent's Role in Advancing Cutting-Edge Therapies, Precision Medicine, and Sustainable Solutions for Global Health Challenges" suggests an exploration of Europe's…
Biotech Breakthroughs in Europe: Exploring the Continent’s Leading Innovations, Research Initiatives, and Collaborative Efforts Shaping the Future of Healthcare and Life Sciences
"Biotech Breakthroughs in Europe: Exploring the Continent's Leading Innovations, Research Initiatives, and Collaborative Efforts Shaping the Future of Healthcare and Life Sciences" suggests a comprehensive…
Ethical Considerations in European Biotech: Balancing Scientific Progress with Societal Values, Human Rights, and Environmental Responsibility in the Pursuit of Innovation
"Ethical Considerations in European Biotech: Balancing Scientific Progress with Societal Values, Human Rights, and Environmental Responsibility in the Pursuit of Innovation" suggests a thorough exploration…
Biotech for All: Europe’s Inclusive Approach to Promoting Diversity, Equity, and Accessibility in Biotechnology Research, Education, and Entrepreneurship
"Biotech for All: Europe's Inclusive Approach to Promoting Diversity, Equity, and Accessibility in Biotechnology Research, Education, and Entrepreneurship" suggests a comprehensive examination of how Europe…
5G Network
Sustainable Smart Cities: 5G’s Role in Building Eco-Friendly Urban Centers in the Middle East
5G technology plays a crucial role in building sustainable smart cities in the Middle East, where the need for efficient resource management and environmental conservation…
Tourism Transformed: Enhancing the Visitor Experience in the Middle East with 5G
The deployment of 5G technology in the Middle East is transforming the tourism industry and enhancing the visitor experience in the region. With its high-speed…
Innovation Hubs: How Middle Eastern Cities Are Pioneering 5G Research and Development
Middle Eastern cities are embracing the potential of 5G technology and positioning themselves as innovation hubs for research and development in the region. Recognizing the…
Connecting the Unconnected: Bridging the Digital Divide with 5G in the Middle East
Bridging the digital divide is a significant challenge for many regions, including the Middle East. The deployment of 5G technology presents a unique opportunity to…
Cultural and Social Transformation: 5G’s Influence on Middle Eastern Communities
The deployment of 5G technology is ushering in a cultural and social transformation in Middle Eastern communities, bringing about significant changes in how people interact,…
5G and Economic Growth: A Middle Eastern Renaissance
The deployment of 5G technology in the Middle East has the potential to catalyze a new era of economic growth and innovation, driving a renaissance…
Security and Privacy in the Age of 5G: Middle East Perspectives
Security and privacy are critical concerns in the age of 5G, and the Middle East is no exception. While 5G technology offers numerous benefits, its…
Navigating Challenges: Implementing 5G Infrastructure in the Middle East
Implementing 5G infrastructure in the Middle East comes with its own set of challenges, ranging from technical and regulatory issues to economic and societal considerations.…
AEROSPACE & DEFENSE
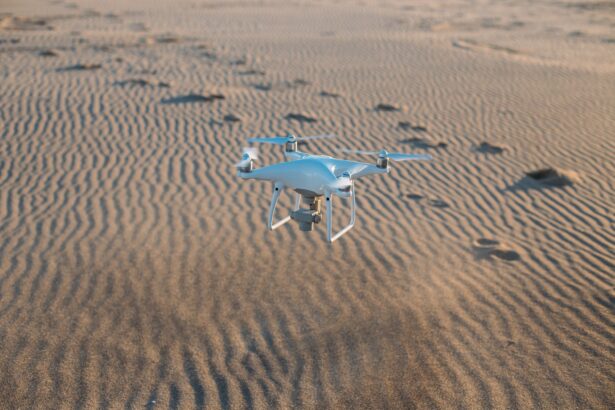
Unmanned Aerial Systems (UAS) Proliferation in Arab Defense
The proliferation of Unmanned Aerial Systems (UAS), commonly known as drones, in Arab defense has been notable in recent years. Arab countries have recognized the…
Arab Military Modernization: The Role of Cutting-Edge Defense Technology
Arab military modernization has been driven by the growing need to address evolving security challenges in the region and bolster defense capabilities. Cutting-edge defense technology…
The Future of Space Tourism: Middle Eastern Companies Venturing into the Cosmos
The future of space tourism holds exciting possibilities, and Middle Eastern companies are starting to venture into the cosmos to offer commercial space travel experiences.…
Space Technology Ambitions: Middle East’s Emerging Space Programs and Satellite Ventures
The Middle East has been actively pursuing ambitious space technology programs and satellite ventures, reflecting the region's growing interest in space exploration, scientific research, and…
Drones for Precision Agriculture: Middle Eastern Farmers Adopting Aerial Farming Tech
Drones, also known as unmanned aerial vehicles (UAVs), are transforming precision agriculture practices in the Middle East. As farmers in the region face challenges related…
Astro-Tech: Middle Eastern Space Startups and Their Contributions to Space Exploration
Astro-Tech, referring to the intersection of space technology and startups, is a burgeoning field in the Middle East. The region has seen the emergence of…
Aerospace Industry Sustainability: Middle Eastern Companies Pursuing Greener Aviation
The aerospace industry is making concerted efforts to become more sustainable and environmentally friendly, and Middle Eastern companies are no exception. With a focus on…
Advancements in Aerospace Technology: Middle Eastern Contributions to Aviation
The Middle East has made significant contributions to aerospace technology and aviation, playing a crucial role in the advancement of the global aerospace industry. Over…
AUTOMOTIVE
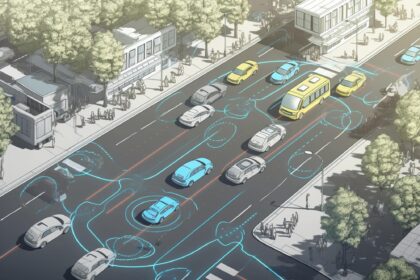
Autonomous Vehicles in Arab Landscapes: Challenges and Opportunities
Autonomous vehicles (AVs) have the potential to revolutionize transportation in Arab landscapes, offering improved safety, efficiency, and convenience. However, their adoption in the region comes…
Electric Vehicle Revolution in the Arab World: Accelerating Sustainability on the Roads
The electric vehicle (EV) revolution is gaining momentum in the Arab world, as countries in the region embrace sustainable transportation solutions to combat climate change,…
Smart Mobility Solutions: Transforming Urban Transport in Arab Cities
Smart mobility solutions are revolutionizing urban transportation in Arab cities, addressing the challenges of traffic congestion, air pollution, and the need for more efficient and…
Connected Car Revolution: IoT’s Impact on Arab Automotive Industry
The connected car revolution is transforming the Arab automotive industry with the widespread adoption of Internet of Things (IoT) technologies in vehicles. IoT integration in…
The Future of Mobility: Electric and Autonomous Vehicles in Middle Eastern Urban Centers
The future of mobility in Middle Eastern urban centers is set to be shaped by electric and autonomous vehicles, with these technologies offering transformative potential…
Sustainable Transportation: Middle Eastern Cities Leading the Shift to Electric Mobility
Middle Eastern cities are at the forefront of the shift to electric mobility, leading the way in adopting sustainable transportation solutions and promoting electric vehicles…
Sustainable Transportation Infrastructure: Middle Eastern Cities Prioritizing Eco-Friendly Projects
Middle Eastern cities are increasingly prioritizing sustainable transportation infrastructure projects to address environmental concerns, reduce carbon emissions, and enhance overall urban livability. The region's commitment…
Smart Transportation Systems: Middle Eastern Cities Navigating Urban Mobility Challenges
Middle Eastern cities are actively embracing smart transportation systems to address the urban mobility challenges posed by rapid urbanization, population growth, and increasing congestion. Smart…
Next-Gen Transportation Infrastructure: Middle East’s Investment in Smart Mobility
The Middle East is investing heavily in next-generation transportation infrastructure and smart mobility solutions to meet the challenges of rapid urbanization, population growth, and the…
Revolutionizing Transportation: How Middle Eastern Cities are Adopting Electric Vehicles
Middle Eastern cities are actively adopting electric vehicles (EVs) as part of their efforts to revolutionize transportation and promote sustainable mobility. The region's commitment to…
Tesla also affected due to corona virus
Tesla, one of the biggest and the most renowned name in the technology sector with great innovations transforming the world was latest in talks due…
Technologies Used In Automobiles
Electronic technology in cars is becoming as crucial as what is under the hood these days. From security and safety to communication and connectivity these…
BIG DATA
Ways Big Data is Revolutionizing Business Operations
Big Data refers to the large and complex data sets that can be analyzed to reveal patterns, trends, and associations. In recent years, Big Data…
Fraud detection with big data analysis
World is moving to the digitized platforms and this digital approach is also giving a strong way to the intruders to enter any digital network…
Big data revolutionizing the marketing industry
Big data is a hot topic in the world of marketing. When you talk about marketing, the sector reflects a lot of challenges and fewer…
Gain Business Enhancements with Big Data
With the requirement of business enhancements along with improved customer experience, every business, governmental institutions have started utilizing the power of Big Data. Big Data…
An introduction to big data
Data storage is really crucial; either you talk about a business or an individual. When it comes to store a huge amount of data for…
Big data: A strong weapon against rising cyber security attacks
The world has went digitized where every single requirements is been satisfied with few finger touches on their mobile screen or other digital screens. Digital…
Wonders of big data in healthcare sector
Data can work wonders in any business sector if processed and mined properly. It has been playing a crucial role in improving services of various…
In what ways big data abetting the digital transformation?
One of the key factor for digital transformation is the big data. An organisation has huge data which is generated, collected frequently so these organisations…
BIOTECH
The European Bioeconomy: A Roadmap to Innovation, Competitiveness, and Sustainability – Policies, Investments, and Partnerships Driving Biotechnology Advancements and Economic Growth
"The European Bioeconomy: A Roadmap to Innovation, Competitiveness, and Sustainability - Policies, Investments, and Partnerships Driving Biotechnology Advancements and Economic Growth" suggests an exploration of…
Investing in Europe’s Biotech Future: Analyzing Funding Trends, Venture Capital Landscape, and Growth Opportunities in the Continent’s Dynamic Life Sciences Industry
"Investing in Europe's Biotech Future: Analyzing Funding Trends, Venture Capital Landscape, and Growth Opportunities in the Continent's Dynamic Life Sciences Industry" suggests an in-depth analysis…
Sustainable Biotech Solutions: Europe’s Contribution to Addressing Environmental Challenges, Sustainable Agriculture, and Circular Economy Principles Through Biotechnology Innovations
"Sustainable Biotech Solutions: Europe's Contribution to Addressing Environmental Challenges, Sustainable Agriculture, and Circular Economy Principles Through Biotechnology Innovations" suggests an exploration of Europe's efforts to…
Revolutionizing Healthcare: How Europe’s Biotech Sector is Redefining Diagnostics, Therapeutics, and Disease Prevention Through Innovations in Molecular Biology and Bioinformatics
"Revolutionizing Healthcare: How Europe's Biotech Sector is Redefining Diagnostics, Therapeutics, and Disease Prevention Through Innovations in Molecular Biology and Bioinformatics" suggests a comprehensive exploration of…
Biosecurity in Europe: Safeguarding Public Health, Biodiversity, and Food Systems – Strategies, Technologies, and Policy Imperatives in the Face of Emerging Biological Threats
"Biosecurity in Europe: Safeguarding Public Health, Biodiversity, and Food Systems - Strategies, Technologies, and Policy Imperatives in the Face of Emerging Biological Threats" suggests a…
Biotech Hubs of Europe: Spotlight on Clusters, Startups, and Research Institutions Driving Innovation and Collaboration in the Continent’s Thriving Life Sciences Ecosystem
"Biotech Hubs of Europe: Spotlight on Clusters, Startups, and Research Institutions Driving Innovation and Collaboration in the Continent's Thriving Life Sciences Ecosystem" suggests an in-depth…
Biotech Resilience in Europe: Navigating Regulatory Frameworks, Investment Trends, and Global Competitiveness in an Era of Scientific Advancement and Economic Uncertainty
"Biotech Resilience in Europe: Navigating Regulatory Frameworks, Investment Trends, and Global Competitiveness in an Era of Scientific Advancement and Economic Uncertainty" suggests an in-depth examination…
Genetic Frontiers: Europe’s Journey in Biotechnology – From CRISPR to Gene Therapy, A Comprehensive Analysis of Innovations, Regulations, and Ethical Considerations
"Genetic Frontiers: Europe's Journey in Biotechnology - From CRISPR to Gene Therapy, A Comprehensive Analysis of Innovations, Regulations, and Ethical Considerations" suggests a detailed exploration…
BLOCKCHAIN
Forging the European Blockchain Agenda: Policy Priorities, Regulatory Frameworks, and Investment Strategies Propelling the Continent Towards Technological Leadership and Global Competitiveness
Forging the European Blockchain Agenda entails setting policy priorities, establishing regulatory frameworks, and implementing investment strategies to propel the continent towards technological leadership and global…
Bridging Borders: The European Blockchain Corridor – An Examination of Cross-Border Collaborations, Interoperability, and Standards Driving Regional Integration and Innovation
Bridging Borders: The European Blockchain Corridor" exemplifies the region's commitment to cross-border collaborations, interoperability, and standards driving regional integration and innovation. Here's a closer look…
Distributed Ledger Diplomacy: European Union’s Strategic Imperatives and Collaborative Efforts in Harnessing Blockchain Technology for Economic Resilience and Technological Sovereignty
Distributed Ledger Diplomacy" aptly describes the European Union's strategic imperatives and collaborative efforts in harnessing blockchain technology for economic resilience and technological sovereignty. Let's delve…
Elevating Trust: How Blockchain Technology is Revolutionizing European Industries, Governance, and Societal Systems for a More Transparent and Inclusive Future
Blockchain technology is revolutionizing European industries, governance, and societal systems, ushering in a new era of transparency, efficiency, and inclusivity. Here's how blockchain is elevating…
Europe’s Blockchain Renaissance: Traversing the Intersection of Technology, Regulation, and Innovation to Shape the Continent’s Digital Destiny
Europe's blockchain renaissance signifies a pivotal moment at the intersection of technology, regulation, and innovation, shaping the continent's digital destiny. Let's explore how these elements…
Charting the Course: European Governments’ Policies, Investments, and Collaborations in Blockchain Technology to Foster Economic Growth, Innovation, and Digital Transformation
European governments are actively charting a course in blockchain technology to foster economic growth, drive innovation, and facilitate digital transformation across various sectors. Here's an…
Blockchain for Social Good: European Initiatives Harnessing Distributed Ledger Technology to Address Environmental, Social, and Governance Challenges in the Region
Blockchain technology is increasingly being leveraged for social good initiatives in Europe, addressing a wide range of environmental, social, and governance (ESG) challenges. These initiatives…
Securing Europe’s Digital Future: Exploring the Role of Blockchain in Safeguarding Data Privacy, Cybersecurity, and Digital Identity Across the Continent
Securing Europe's digital future is a paramount concern in an increasingly interconnected and data-driven world. Blockchain technology holds significant promise in safeguarding data privacy, enhancing…
Blockchain Strategies for European Enterprises: Harnessing the Power of Distributed Ledger Technology to Drive Innovation, Efficiency, and Competitiveness in the Global Market
European enterprises are increasingly recognizing the transformative potential of blockchain technology in driving innovation, efficiency, and competitiveness in the global market. To develop effective blockchain…
The European Blockchain Odyssey: Navigating the Complexities of Adoption, Integration, and Regulation in a Continent of Diverse Cultures and Economies
The European Blockchain Odyssey presents a multifaceted journey through the complexities of adoption, integration, and regulation across a continent characterized by diverse cultures and economies.…
European Blockchain Renaissance: A Comprehensive Overview of Emerging Trends, Promising Use Cases, and Regulatory Considerations Shaping the Continent’s Blockchain Landscape
The European blockchain landscape is experiencing a renaissance, marked by a surge in innovation, adoption, and regulatory developments. Let's delve into the key trends, promising…
Trust in the Digital Age: How European Nations are Leveraging Blockchain to Build Transparent, Secure, and Inclusive Systems for the Future
In recent years, European nations have increasingly turned to blockchain technology to address various challenges related to trust, transparency, security, and inclusivity. Blockchain, a decentralized…
CLOUD COMPUTING
E-commerce Expansion: Cloud Solutions Driving Online Retail Growth in the Middle East
Cloud solutions are playing a significant role in driving the expansion and growth of e-commerce in the Middle East. As the region's online retail market…
Cloud Skills Development: Bridging the Talent Gap in the Middle East
Cloud skills development is crucial in bridging the talent gap in the Middle East and meeting the increasing demand for skilled professionals in cloud computing.…
Hybrid Cloud Solutions: Balancing On-Premises and Cloud Infrastructure in the Middle East
Hybrid cloud solutions have gained popularity in the Middle East as they offer a flexible and balanced approach that combines on-premises infrastructure with cloud services.…
Building Resilience: Cloud Disaster Recovery Strategies for Middle Eastern Businesses
Building resilience through cloud disaster recovery strategies is crucial for Middle Eastern businesses to protect their critical data and services from potential disruptions. Cloud-based disaster…
Data Security in the Cloud: Middle Eastern Approaches and Concerns
Data security in the cloud is a critical aspect of cloud adoption in the Middle East, as organizations seek to protect sensitive information while leveraging…
Empowering Startups: How Cloud Computing is Fueling Innovation in the Middle East
Cloud computing has become a powerful catalyst for innovation in the Middle East's startup ecosystem, empowering entrepreneurs to develop and scale their ventures more efficiently.…
Cloud Adoption in the Middle East: Trends and Transformations
Cloud adoption in the Middle East has been steadily growing over the years, and it has undergone significant transformations, driven by several trends and factors…
Cloud-Native Middle East: Accelerating Innovation with Serverless and Containers
The cloud-native approach, leveraging serverless computing and containers, is gaining momentum in the Middle East as organizations seek to accelerate innovation, improve agility, and enhance…
Sustainability in the Cloud: Green Computing Initiatives in the Middle East
Sustainability and green computing initiatives in the cloud are gaining traction in the Middle East as organizations become more conscious of their environmental impact and…
Cloud technology: The new face of today’s start-up
Startups are growing at the fastest pace bringing astonishing and creative ideas into vitality using the latest technologies. The inception of new ideas requires a…
Advantages of Cloud Computing for business development
Traditional business strategies were much bonded within papers and offered limited access. With the emergence of cloud computing most of the businesses are on boarding…
Accelerate Business growth with Cloud Computing
Cloud Computing is the process of outsourcing of software, data storage, and processing. With the help of an internet connection, users have started accessing applications…
CYBER SECURITY
Cybersecurity Investment Trends in Europe: Analyzing Funding Patterns, Market Dynamics, and Growth Opportunities in the Continent’s Booming Cybersecurity Industry
Cybersecurity Investment Trends in Europe: Analyzing Funding Patterns, Market Dynamics, and Growth Opportunities in the Continent's Booming Cybersecurity Industry" suggests an exploration of the investment…
From Brussels to Berlin: Tracing the Evolution of European Cybersecurity Policies and Regulations – Impact, Implementation, and Future Directions
From Brussels to Berlin: Tracing the Evolution of European Cybersecurity Policies and Regulations - Impact, Implementation, and Future Directions" suggests a detailed examination of how…
The European Cybersecurity Ecosystem: A Comprehensive Analysis of Public-Private Partnerships, Innovation Clusters, and Research Initiatives Driving Technological Advancement and Collaboration
The European Cybersecurity Ecosystem: A Comprehensive Analysis of Public-Private Partnerships, Innovation Clusters, and Research Initiatives Driving Technological Advancement and Collaboration" suggests a thorough examination of…
Cybersecurity as a Competitive Advantage: How European Businesses Are Embracing Proactive Risk Management, Incident Response, and Cyber Defense Strategies
Cybersecurity as a Competitive Advantage: How European Businesses Are Embracing Proactive Risk Management, Incident Response, and Cyber Defense Strategies" suggests an exploration of how European…
Cyber Hygiene and Education: Europe’s Efforts to Build a Cyber-Aware Society – Strategies, Initiatives, and Challenges in Promoting Cybersecurity Awareness and Resilience
Cyber Hygiene and Education: Europe's Efforts to Build a Cyber-Aware Society - Strategies, Initiatives, and Challenges in Promoting Cybersecurity Awareness and Resilience" suggests an in-depth…
Cyber Threat Intelligence in Europe: Leveraging Data, Analytics, and Collaboration to Stay Ahead of Sophisticated Adversaries and Emerging Threat Vectors
Cyber Threat Intelligence in Europe: Leveraging Data, Analytics, and Collaboration to Stay Ahead of Sophisticated Adversaries and Emerging Threat Vectors" suggests a deep dive into…
Beyond Compliance: Europe’s Proactive Approach to Cybersecurity – Innovations, Best Practices, and Emerging Technologies for a Secure Digital Ecosystem
Beyond Compliance: Europe's Proactive Approach to Cybersecurity - Innovations, Best Practices, and Emerging Technologies for a Secure Digital Ecosystem" suggests an exploration of how Europe…
Guardians of the Digital Domain: Exploring Europe’s Role in Global Cybersecurity Governance, Threat Intelligence Sharing, and Norm Development
Guardians of the Digital Domain: Exploring Europe's Role in Global Cybersecurity Governance, Threat Intelligence Sharing, and Norm Development" suggests a focused exploration of Europe's contributions…
Cyber Resilience in the European Union: Strengthening Defenses, Fostering Collaboration, and Navigating Regulatory Imperatives in an Era of Persistent Threats
Cyber Resilience in the European Union: Strengthening Defenses, Fostering Collaboration, and Navigating Regulatory Imperatives in an Era of Persistent Threats" encapsulates a thorough investigation into…
Securing Europe’s Digital Future: A Holistic Examination of Cybersecurity Challenges, Strategies, and Innovations Driving Resilience and Trust Across the Continent
Securing Europe's Digital Future: A Holistic Examination of Cybersecurity Challenges, Strategies, and Innovations Driving Resilience and Trust Across the Continent" suggests a comprehensive exploration of…
Safeguarding the European Cyberspace: A Deep Dive into the Threat Landscape, Defense Mechanisms, and Future Directions of Cybersecurity in the Continent
Safeguarding the European Cyberspace: A Deep Dive into the Threat Landscape, Defense Mechanisms, and Future Directions of Cybersecurity in the Continent" is a title that…
Beyond Borders: European Union’s Unified Front Against Cyber Threats – Strategies, Partnerships, and Policies for Ensuring Digital Sovereignty and Resilience
Beyond Borders: European Union's Unified Front Against Cyber Threats - Strategies, Partnerships, and Policies for Ensuring Digital Sovereignty and Resilience" is a title that suggests…
AR & VR
The Future of Augmented Reality in Travel: Enhancing Middle Eastern Tourism
The future of augmented reality (AR) in travel holds immense potential for enhancing tourism experiences in the Middle East. As AR technology continues to advance,…
Virtual Design for Middle Eastern Festivals: Immersive Celebrations in the Digital Age
Virtual design is redefining the way Middle Eastern festivals are celebrated, offering immersive and interactive experiences in the digital age. With the advent of virtual…
Augmented Reality in Tourism: Enhancing Middle Eastern Cultural Experiences
Augmented Reality (AR) is revolutionizing tourism in the Middle East by enhancing cultural experiences and offering visitors a deeper understanding of the region's rich heritage.…
Augmented Reality in Heritage Sites: Enhancing Middle Eastern Tourist Experiences
Augmented Reality (AR) is playing a transformative role in enhancing tourist experiences at heritage sites in the Middle East. AR technology overlays digital information, images,…
Virtual Reality Training: Preparing Middle Eastern Workforces for High-Risk Industries
Virtual Reality (VR) training is becoming an increasingly valuable tool in preparing Middle Eastern workforces for high-risk industries. VR simulations offer a safe and controlled…
Virtual Reality Therapy: Middle Eastern Applications for Mental Health Treatment
Virtual Reality Therapy (VRT) is gaining traction as an innovative and effective approach to mental health treatment in the Middle East. VRT uses virtual reality…
The Role of Virtual Reality in Architecture: Redefining Urban Planning in the Middle East
Virtual Reality (VR) is redefining urban planning in the Middle East by offering architects, urban planners, and policymakers new tools and perspectives to envision and…
The Role of Virtual Reality in Architecture: Middle Eastern Designs of the Future
Virtual Reality (VR) is playing a transformative role in the field of architecture in the Middle East, revolutionizing the design and visualization process of future…
Immersive Virtual Events: How Middle Eastern Conferences are Embracing Virtual Reality
Middle Eastern conferences are increasingly embracing immersive virtual events, leveraging virtual reality (VR) technology to host interactive and engaging gatherings. These immersive virtual events offer…
Exploring AR/VR in Education: Immersive Learning Experiences in Middle Eastern Schools
AR/VR (Augmented Reality/Virtual Reality) technologies are transforming the education landscape in Middle Eastern schools, offering immersive and interactive learning experiences to students. These cutting-edge technologies…
What are the advanced applications of virtual reality?
Virtual reality is gaining a rapid popularity among folks due to its amazing features that let you penetrate into a virtual world and experience it…
Miracles of virtual reality in physical therapy
Every year, there are more than a billion people struggle from strokes and traumatic brain injuries. These injuries are curable and require physical therapies to…
EDUCATION
AI-Powered Personalized Learning: Transforming Education in Middle Eastern Schools
AI-powered personalized learning is revolutionizing education in Middle Eastern schools, providing tailored and adaptive learning experiences to students based on their individual needs, abilities, and…
Virtual Reality in Religious Education: Immersive Learning in Middle Eastern Seminaries
Virtual Reality (VR) is revolutionizing religious education in Middle Eastern seminaries by offering immersive and experiential learning experiences for students and scholars. VR technology allows…
Virtual Museums and Cultural Diplomacy: Middle Eastern Heritage in the Digital Age
Virtual museums and cultural diplomacy are playing a vital role in preserving and promoting Middle Eastern heritage in the digital age. These initiatives leverage technology…
The Role of Robotics in Middle Eastern Education: Transforming Learning and Teaching
The role of robotics in Middle Eastern education is transforming the learning and teaching landscape, as educators and policymakers recognize the potential of robotics to…
The Intersection of Gaming and Education: Gamification in Middle Eastern Curricula
The intersection of gaming and education, often referred to as gamification, is gaining traction in Middle Eastern curricula as educators recognize the potential of using…
The Future of Quantum Computing in Middle Eastern Research Institutions
The future of quantum computing in Middle Eastern research institutions holds great promise for advancing scientific discoveries, solving complex problems, and revolutionizing various industries. Quantum…
Tech for Inclusive Education: Assisting Students with Disabilities in the Middle East
Tech for inclusive education is playing a pivotal role in assisting students with disabilities in the Middle East, ensuring they have equal access to quality…
Rise of E-learning: Technology’s Impact on Education during Challenging Times in the Middle East
The rise of e-learning has been instrumental in transforming education during challenging times in the Middle East, particularly in the face of crises like the…
Rise of Edutainment: Middle Eastern Startups Combining Education and Entertainment
The rise of edutainment, the fusion of education and entertainment, is gaining momentum in the Middle East as startups recognize the potential of using interactive…
Inclusive Tech Gaming: Middle Eastern Developers Creating Accessible Video Games
Inclusive tech gaming is gaining traction in the Middle East, as developers recognize the importance of creating accessible video games that cater to a diverse…
Tips for developing an inclusive education system
In today’s era, where everyone is deeming for human rights equality are highly working for the equality of genders, cast, creed, and other races. “What…
How to use technology in Education?
Technology is revolutionizing the complete world in a better way. It has transformed education also, by empowering it by giving them the ownership of how…
FINTECH
Insurtech Illumination: Shedding Light on the Role of Artificial Intelligence and Big Data Analytics in European Insurance Innovation
Insurtech Illumination: Shedding Light on the Role of Artificial Intelligence and Big Data Analytics in European Insurance Innovation" suggests a focused exploration of how artificial…
Beyond Traditional Coverage: The Rise of Insurtech Startups and Their Disruptive Influence on Europe’s Insurance Industry
Beyond Traditional Coverage: The Rise of Insurtech Startups and Their Disruptive Influence on Europe's Insurance Industry" suggests an exploration of how insurtech startups are challenging…
EuroInsights: Uncovering the Latest Trends and Insights Driving Fintech Growth in Europe
EuroInsights: Uncovering the Latest Trends and Insights Driving Fintech Growth in Europe" suggests an in-depth exploration of the current trends and insights shaping the fintech…
Beyond Banking: Exploring the Diverse Applications of Fintech in Europe
Beyond Banking: Exploring the Diverse Applications of Fintech in Europe" suggests an exploration of how fintech innovations extend beyond traditional banking services and into various…
EuroFintech Ecosystem: Mapping the Dynamics of Innovation Across Europe
EuroFintech Ecosystem: Mapping the Dynamics of Innovation Across Europe" suggests an exploration of the diverse landscape of fintech innovation across European countries. Here's how we…
The Eurocoin Revolution: Exploring Europe’s Cryptocurrency Adoption Journey
The Eurocoin Revolution: Exploring Europe's Cryptocurrency Adoption Journey" suggests an exploration of how cryptocurrencies, particularly Eurocoin, are being adopted and integrated into various facets of…
EuroTech Renaissance: Revitalizing Traditional Banking with Fintech in Europe
EuroTech Renaissance: Revitalizing Traditional Banking with Fintech in Europe" suggests an exploration of how fintech innovations are revitalizing and transforming traditional banking practices across Europe.…
From PSD2 to PSD3: The Evolution of European Payment Services Directive and Its Impact on Fintech
From PSD2 to PSD3: The Evolution of European Payment Services Directive and Its Impact on Fintech" suggests an exploration of how the Payment Services Directive…
EuroPay: The Evolution of Payment Systems in Europe’s Digital Economy
EuroPay: The Evolution of Payment Systems in Europe's Digital Economy" suggests an exploration of the transformation of payment systems within Europe's digital landscape. Here's a…
Fintech Frontiers: Navigating the Regulatory Landscape in Europe’s Financial Sector
Fintech Frontiers: Navigating the Regulatory Landscape in Europe's Financial Sector" suggests an exploration of the regulatory challenges and opportunities facing fintech companies operating in Europe's…
Smart Contracts, Smarter Europe: The Impact of Blockchain on European Business
Smart Contracts, Smarter Europe: The Impact of Blockchain on European Business" suggests an examination of how blockchain technology and smart contracts are influencing business practices…
Financial Inclusion Revolution: Bridging the Gap Through Fintech in Europe
Financial Inclusion Revolution: Bridging the Gap Through Fintech in Europe" suggests an exploration of how fintech innovations are helping to address the issue of financial…
GOVERNMENT & PUBLIC SECTOR
Middle Eastern Governments Embrace AI: Transforming Public Services and Decision-Making
Middle Eastern governments are increasingly embracing Artificial Intelligence (AI) to transform public services and decision-making processes. AI adoption in the region is driven by the…
E-Government Evolution in the Middle East: Digital Transformation for Seamless Citizen Engagement
The evolution of e-government in the Middle East has seen significant strides over the years, as governments in the region embrace digital transformation to provide…
Blockchain in Middle Eastern Government: Ensuring Transparency and Trust in Public Transactions
Blockchain technology is being increasingly adopted by Middle Eastern governments to enhance transparency, security, and trust in public transactions. As a decentralized and immutable ledger…
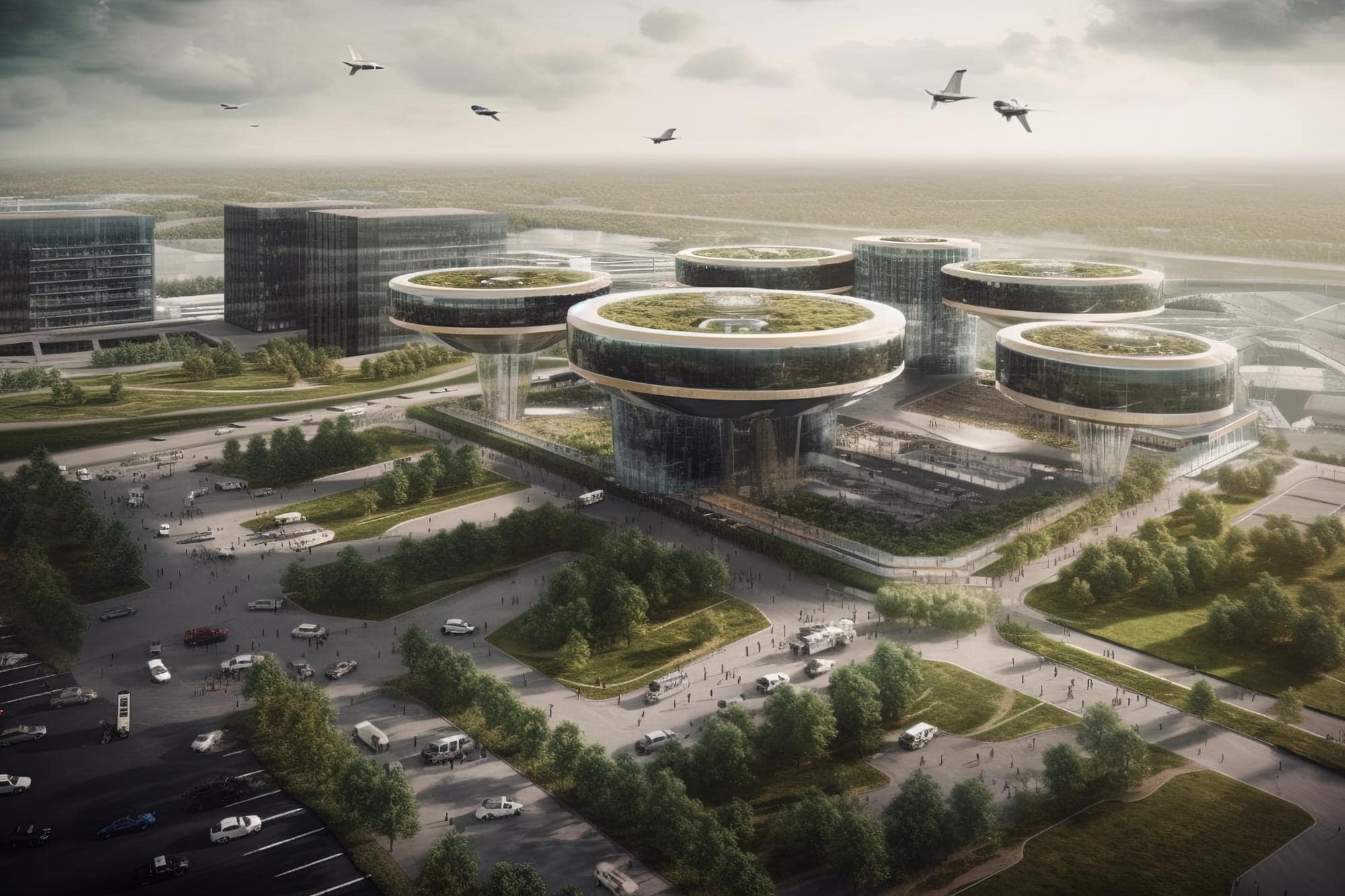
Smart Cities in the Middle East: Advancing Urban Governance Through Technology
Smart cities in the Middle East are at the forefront of urban governance advancements, harnessing technology to improve city services, enhance efficiency, and create a…
Cultural Heritage Preservation: Government Efforts to Safeguard History and Identity in the Middle East
Cultural heritage preservation is of paramount importance in the Middle East, given the region's rich history and diverse cultural identity. Middle Eastern governments recognize the…
Innovation Hubs and Smart Cities: Middle Eastern Governments Fostering Technological Advancements
Middle Eastern governments are actively fostering technological advancements through innovation hubs and smart city initiatives. These initiatives aim to promote innovation, attract talent, drive economic…
Youth Empowerment and Middle Eastern Governance: Engaging the Next Generation
Youth empowerment is a critical aspect of governance in the Middle East, as the region has a significant youth population with diverse aspirations and potential.…
Sustainable Development Goals in the Middle East: Government Initiatives and Progress
The Sustainable Development Goals (SDGs) are a set of global goals adopted by the United Nations to address various social, economic, and environmental challenges by…
Middle Eastern Governments’ Response to Crisis: Navigating Challenges and Ensuring Resilience
Middle Eastern governments have faced various crises over the years, including economic challenges, geopolitical tensions, natural disasters, and the COVID-19 pandemic. In response to these…
Cybersecurity Imperatives for Middle Eastern Governments: Protecting Critical Infrastructure and Data
Cybersecurity is a critical imperative for Middle Eastern governments as they strive to protect their critical infrastructure and sensitive data from cyber threats. The region…
Communicating legit information in this pandemic
The COVID-19 (coronavirus) is plausibly the pandemic that has worsen the situation in the whole world. Since its inception to date, there have been various…
Modernization by technology
An efficacious government is a cornerstone for developing the country. Also, upheave in citizens expectation has made the government perform well. The potential to meet…
Efficient government technologies
We are in the era where a blanket of technologies is being unfolded, that is being used in different ways in daily lives, on which…
Top GovTech Trends Revealed for 2021
The year 2021 has come up with a wide array of changes for the entire world. The government is also moving up front to bring…
Major challenges faced by army
Army constitutes a major part of government sector has also plays a crucial role in saving an entire country by protecting its borders from the…
Strategies to Improve Public Sector Labour Productivity
Labour productive is an essential element for any organization to perform well and achieve intended goal. To keep your organizational working procedure effective and accurate,…
ENERGY AND UTILITIES
Water Technology Advancements in the Middle East: Desalination, Purification, and Resource Management
Water technology advancements in the Middle East are critical in addressing the region's water scarcity and ensuring sustainable water resource management. The region faces arid…
Renewable Energy Integration in the Middle East: Tackling Grid Stability and Storage Challenges
Renewable energy integration in the Middle East presents both opportunities and challenges, particularly concerning grid stability and energy storage. As the region increasingly adopts renewable…
Smart Metering and Energy Efficiency: Overcoming Adoption Challenges in Middle East Utilities
Smart metering and energy efficiency initiatives are crucial components of the energy transition in the Middle East. However, the adoption of smart metering faces some…
Carbon Emission Reduction Strategies in the Middle East: Transitioning Towards Net-Zero Goals
Carbon emission reduction strategies are becoming increasingly important in the Middle East as countries in the region recognize the urgency of addressing climate change and…
Decentralized Energy Generation: Microgrids and Distributed Energy Resources in the Middle East
Decentralized energy generation, particularly through microgrids and distributed energy resources (DERs), is emerging as a viable solution to address energy challenges in the Middle East.…
Water Scarcity and Sustainable Utilities: Innovations Addressing Middle East’s Growing Water Crisis
Water scarcity is a pressing issue in the Middle East, where arid and semi-arid climates pose significant challenges to water availability. Sustainable water utilities and…
Smart Grids and Energy Management: Overcoming Grid Integration Challenges in the Middle East
Smart grids and advanced energy management systems are essential components of the energy transition in the Middle East. However, integrating smart grids into existing energy…
Energy Transition in the Middle East: Balancing Renewable Integration and Fossil Fuel Dependence
The energy transition in the Middle East is a complex process that involves balancing the integration of renewable energy sources with the region's historical dependence…
HEALTHCARE
Virtual Medical Tourism: Middle Eastern Hospitals Attracting International Patients Online
Virtual medical tourism, also known as telemedical tourism or remote medical tourism, is an emerging trend in the healthcare industry, and Middle Eastern hospitals are…
The Intersection of Biotechnology and Agriculture: Innovations in Middle Eastern Farming
The intersection of biotechnology and agriculture has the potential to revolutionize farming practices in the Middle East, addressing challenges related to food security, water scarcity,…
The Impact of 5G on Healthcare: Middle Eastern Hospitals Embracing Next-Gen Connectivity
The deployment of 5G technology in healthcare is set to revolutionize the industry, and Middle Eastern hospitals are increasingly embracing the next-generation connectivity to improve…
The Impact of 3D Printing in Healthcare: Middle Eastern Hospitals Embracing Additive Manufacturing
3D printing, also known as additive manufacturing, has had a significant impact on the healthcare industry globally, and Middle Eastern hospitals are increasingly embracing this…
The Evolution of Mobile Health Apps: Empowering Healthcare Consumers in the Middle East
The evolution of mobile health apps has been transformative, especially in empowering healthcare consumers in the Middle East. These apps, commonly known as mHealth apps,…
Tech-Enhanced Traditional Medicine: Integrating Ancient Healing with Modern Tools
Integrating modern technology with traditional medicine, often referred to as tech-enhanced traditional medicine or digital traditional medicine, is an innovative approach that leverages the benefits…
Tech-enabled Healthcare: Telemedicine and Digital Health Services in the Middle East
the adoption of tech-enabled healthcare, including telemedicine and digital health services, was gaining momentum globally, including in the Middle East. However, specific developments may have…
RehabTech: Middle Eastern Innovations Revolutionizing Rehabilitation and Assistive Devices
RehabTech, short for Rehabilitation Technology, refers to innovations and technologies that aim to enhance the rehabilitation process and improve the quality of life for individuals…
MedTech for Bedouin Communities: Tailored Healthcare Solutions in Remote Areas
MedTech, or Medical Technology, has the potential to revolutionize healthcare delivery in remote and underserved areas, including Bedouin communities. Bedouin communities often reside in remote…
Precision Medicine in the Middle East: Tailoring Healthcare with Genomics and AI
Precision medicine, also known as personalized medicine, is an innovative approach to healthcare that considers individual variability in genes, environment, and lifestyle. Integrating genomics and…
How robots can make surgery more efficient?
Surgical robots are not a new name in the medical field. It has been serving the industry for years. But what’s new about it? It’s…
Harness the power of technology in the medical sector for senior citizens
A common life needs medical assistance in many ways to lead a healthy life. But the rising cost of medical solutions for several life-threatening diseases…
Potential of nanotechnology in medical sector
The word nano take us to the tiniest shapes but the measurement of nano is much smaller than what we expect. Nano can be defined…
Healthcare and technology moving hand on hand for the best results
Advancements in the medical sector is not hidden from anyone, every single eye in this world has witnessed a great transformation in the medical assistance…
Impact of technology on health care sector
A new era of telemedicine has been emerged with the advancements of technology and its use in the healthcare industry. Evolution of technology has enabled…
A complete guide on healthcare technology and its uses
Healthcare sector is a boon for the mankind that assists folks to regain their healthy and live it to their fullest. It is a digitized…
HR TECHNOLOGY
Diversity and Inclusion Tech Solutions in the Middle East Fostering a More Equitable Workplace
Diversity and inclusion tech solutions in the Middle East are playing a crucial role in fostering a more equitable and inclusive workplace environment. These solutions…
Remote Work Revolution in the Middle East HR Tech’s Role in Shaping the Future of Work
The remote work revolution in the Middle East has been accelerated by the COVID-19 pandemic and the increasing adoption of digital technologies. As more organizations…
Learning and Development Tech Upskilling the Middle Eastern Workforce for the Digital Age
Learning and development tech is playing a crucial role in upskilling the Middle Eastern workforce for the digital age. As the region embraces digital transformation…
HR Tech Startups in the Middle East Innovations and Disruptions in the HR Landscape
HR tech startups in the Middle East are driving innovations and disruptions in the HR landscape by introducing new technologies, tools, and solutions to enhance…
HR Analytics and Data-Driven Insights Making Informed Decisions in Middle Eastern Organizations
HR analytics and data-driven insights are playing a crucial role in helping Middle Eastern organizations make informed and strategic decisions about their workforce. By leveraging…
Gig Economy and Freelancer Platforms in the Middle East Reshaping Talent Management
The gig economy and freelancer platforms are reshaping talent management in the Middle East, providing both opportunities and challenges for businesses in the region. As…
Ethical AI in HR Ensuring Fairness and Bias-Free Decisions in the Middle East
Ethical AI in HR is critical to ensuring fairness, transparency, and bias-free decision-making in the Middle Eastern workplace. As AI and machine learning algorithms play…
Employee Wellbeing Tech Prioritizing Wellness in the Middle Eastern Workplace
Employee well-being tech solutions are becoming increasingly important in the Middle Eastern workplace as organizations recognize the significance of prioritizing employee wellness for overall productivity,…
INSURTECH
From Paperwork to Platforms: The Digital Transformation of Claims Processing and Customer Engagement in Europe’s Insurtech Revolution
From Paperwork to Platforms: The Digital Transformation of Claims Processing and Customer Engagement in Europe's Insurtech Revolution" suggests an in-depth exploration of how digital platforms…
Insurtech Interconnect: Navigating Partnerships and Collaborations to Accelerate Innovation and Market Penetration Across Europe
Insurtech Interconnect: Navigating Partnerships and Collaborations to Accelerate Innovation and Market Penetration Across Europe" suggests a comprehensive exploration of how partnerships and collaborations can drive…
Reshaping Risk Assessment: Leveraging Machine Learning and Predictive Analytics to Revolutionize Underwriting Practices in European Insurtech
Reshaping Risk Assessment: Leveraging Machine Learning and Predictive Analytics to Revolutionize Underwriting Practices in European Insurtech" suggests an exploration of how machine learning (ML) and…
Insurtech Insights: Unlocking the Potential of Telematics, IoT, and Data Analytics to Drive Insurance Innovation in Europe
Insurtech Insights: Unlocking the Potential of Telematics, IoT, and Data Analytics to Drive Insurance Innovation in Europe" suggests a deep dive into how technologies such…
The Insurtech Imperative: Strategies for Incumbents and New Entrants to Thrive in Europe’s Evolving Insurance Landscape
The Insurtech Imperative: Strategies for Incumbents and New Entrants to Thrive in Europe's Evolving Insurance Landscape" suggests an in-depth examination of strategies that both established…
Innovating for Resilience: Harnessing Insurtech Innovations to Address Key Challenges Facing European Insurance Markets
Innovating for Resilience: Harnessing Insurtech Innovations to Address Key Challenges Facing European Insurance Markets" suggests a comprehensive exploration of how insurtech innovations can help tackle…
Insurtech Horizons: A Deep Dive into the Emerging Trends and Opportunities Shaping the Future of Insurance in Europe
Insurtech Horizons: A Deep Dive into the Emerging Trends and Opportunities Shaping the Future of Insurance in Europe" suggests an in-depth exploration of the evolving…
The Digital Insurance Revolution: Exploring Europe’s Journey Towards Seamless, Customer-Centric Insurtech Solutions
The Digital Insurance Revolution: Exploring Europe's Journey Towards Seamless, Customer-Centric Insurtech Solutions" suggests an exploration of how Europe is undergoing a transformation in its insurance…
Tech-Driven Transformation: Examining the Regulatory Landscape and Policy Implications for Insurtech Ventures in Europe
Tech-Driven Transformation: Examining the Regulatory Landscape and Policy Implications for Insurtech Ventures in Europe" suggests an analysis focusing on how the regulatory landscape and policies…
Insurtech Evolution: A Comprehensive Analysis of the Rapid Technological Advancements Transforming the European Insurance Sector
Insurtech Evolution: A Comprehensive Analysis of the Rapid Technological Advancements Transforming the European Insurance Sector" suggests a thorough examination of how technological advancements, particularly within…
Navigating the Digital Insurance Frontier: Unveiling Europe’s Insurtech Ecosystem and Its Impact on Traditional Insurance Models
Navigating the Digital Insurance Frontier: Unveiling Europe's Insurtech Ecosystem and Its Impact on Traditional Insurance Models" suggests an in-depth exploration of how Europe's insurtech ecosystem…
Insurtech Renaissance: Reshaping Insurance Dynamics Across the Eurozone with Innovative Technological Solutions
Insurtech Renaissance: Reshaping Insurance Dynamics Across the Eurozone with Innovative Technological Solutions" suggests an exploration of how insurtech innovations are reshaping the insurance industry dynamics…
Revolutionizing Risk Management: Exploring the Intersection of Technology and Insurance in Europe’s Insurtech Landscape
Revolutionizing Risk Management: Exploring the Intersection of Technology and Insurance in Europe's Insurtech Landscape" suggests an in-depth exploration of how technology is transforming risk management…
The Future of Claims: AI, Drones, and Augmented Reality in Middle Eastern Insurance
The future of claims processing in the Middle Eastern insurance industry is set to be transformed by cutting-edge technologies like AI, drones, and augmented reality.…
Regulatory Landscape for InsurTech: Navigating Compliance Challenges in the Middle East
The regulatory landscape for InsurTech in the Middle East is evolving as the region embraces digital innovation in the insurance industry. InsurTech startups and traditional…
InsurTech Startups in the Middle East: Driving Innovation and Disruption
InsurTech startups are playing a significant role in driving innovation and disruption in the insurance industry across the Middle East. These startups leverage technology, data…
IOT
From Paperwork to Platforms: The Digital Transformation of Claims Processing and Customer Engagement in Europe’s Insurtech Revolution
From Paperwork to Platforms: The Digital Transformation of Claims Processing and Customer Engagement in Europe's Insurtech Revolution" suggests an in-depth exploration of how digital platforms…
Insurtech Interconnect: Navigating Partnerships and Collaborations to Accelerate Innovation and Market Penetration Across Europe
Insurtech Interconnect: Navigating Partnerships and Collaborations to Accelerate Innovation and Market Penetration Across Europe" suggests a comprehensive exploration of how partnerships and collaborations can drive…
Reshaping Risk Assessment: Leveraging Machine Learning and Predictive Analytics to Revolutionize Underwriting Practices in European Insurtech
Reshaping Risk Assessment: Leveraging Machine Learning and Predictive Analytics to Revolutionize Underwriting Practices in European Insurtech" suggests an exploration of how machine learning (ML) and…
Insurtech Insights: Unlocking the Potential of Telematics, IoT, and Data Analytics to Drive Insurance Innovation in Europe
Insurtech Insights: Unlocking the Potential of Telematics, IoT, and Data Analytics to Drive Insurance Innovation in Europe" suggests a deep dive into how technologies such…
The Insurtech Imperative: Strategies for Incumbents and New Entrants to Thrive in Europe’s Evolving Insurance Landscape
The Insurtech Imperative: Strategies for Incumbents and New Entrants to Thrive in Europe's Evolving Insurance Landscape" suggests an in-depth examination of strategies that both established…
Innovating for Resilience: Harnessing Insurtech Innovations to Address Key Challenges Facing European Insurance Markets
Innovating for Resilience: Harnessing Insurtech Innovations to Address Key Challenges Facing European Insurance Markets" suggests a comprehensive exploration of how insurtech innovations can help tackle…
Insurtech Horizons: A Deep Dive into the Emerging Trends and Opportunities Shaping the Future of Insurance in Europe
Insurtech Horizons: A Deep Dive into the Emerging Trends and Opportunities Shaping the Future of Insurance in Europe" suggests an in-depth exploration of the evolving…
The Digital Insurance Revolution: Exploring Europe’s Journey Towards Seamless, Customer-Centric Insurtech Solutions
The Digital Insurance Revolution: Exploring Europe's Journey Towards Seamless, Customer-Centric Insurtech Solutions" suggests an exploration of how Europe is undergoing a transformation in its insurance…
Tech-Driven Transformation: Examining the Regulatory Landscape and Policy Implications for Insurtech Ventures in Europe
Tech-Driven Transformation: Examining the Regulatory Landscape and Policy Implications for Insurtech Ventures in Europe" suggests an analysis focusing on how the regulatory landscape and policies…
Insurtech Evolution: A Comprehensive Analysis of the Rapid Technological Advancements Transforming the European Insurance Sector
Insurtech Evolution: A Comprehensive Analysis of the Rapid Technological Advancements Transforming the European Insurance Sector" suggests a thorough examination of how technological advancements, particularly within…
Navigating the Digital Insurance Frontier: Unveiling Europe’s Insurtech Ecosystem and Its Impact on Traditional Insurance Models
Navigating the Digital Insurance Frontier: Unveiling Europe's Insurtech Ecosystem and Its Impact on Traditional Insurance Models" suggests an in-depth exploration of how Europe's insurtech ecosystem…
Insurtech Renaissance: Reshaping Insurance Dynamics Across the Eurozone with Innovative Technological Solutions
Insurtech Renaissance: Reshaping Insurance Dynamics Across the Eurozone with Innovative Technological Solutions" suggests an exploration of how insurtech innovations are reshaping the insurance industry dynamics…
Revolutionizing Risk Management: Exploring the Intersection of Technology and Insurance in Europe’s Insurtech Landscape
Revolutionizing Risk Management: Exploring the Intersection of Technology and Insurance in Europe's Insurtech Landscape" suggests an in-depth exploration of how technology is transforming risk management…
The Future of Claims: AI, Drones, and Augmented Reality in Middle Eastern Insurance
The future of claims processing in the Middle Eastern insurance industry is set to be transformed by cutting-edge technologies like AI, drones, and augmented reality.…
Regulatory Landscape for InsurTech: Navigating Compliance Challenges in the Middle East
The regulatory landscape for InsurTech in the Middle East is evolving as the region embraces digital innovation in the insurance industry. InsurTech startups and traditional…
InsurTech Startups in the Middle East: Driving Innovation and Disruption
InsurTech startups are playing a significant role in driving innovation and disruption in the insurance industry across the Middle East. These startups leverage technology, data…
LOGISTICS
Smart Ports and Connected Supply Chains: Middle East Logistics Embrace IoT for Efficiency Gains
The Middle East logistics sector is embracing the Internet of Things (IoT) to transform its operations, with a particular focus on smart ports and connected…
Data Integration and Interoperability: Middle East Logistics Tech’s Quest for Unified Systems
Data integration and interoperability are key challenges in the quest for unified systems in Middle East logistics technology. As logistics operations become increasingly digitized, the…
E-Commerce Surge and Middle East Logistics Tech: Innovations to Meet Growing Demand
The e-commerce surge in the Middle East has driven significant innovations in logistics technology to meet the growing demand for efficient and reliable delivery services.…
Digital Transformation of Middle East Logistics: Overcoming Tech Challenges for Seamless Operations
The digital transformation of Middle East logistics is essential for overcoming traditional challenges and achieving seamless and efficient operations. Embracing technology can significantly enhance supply…
Sustainability and Green Logistics in the Middle East: Balancing Economic Growth with Environmental Responsibility
Sustainability and green logistics are becoming increasingly important in the Middle East as the region balances its economic growth with environmental responsibility. The Middle East…
Port and Infrastructure Bottlenecks: Addressing Middle East Logistics Capacity Constraints
Addressing port and infrastructure bottlenecks is crucial to overcoming logistics capacity constraints in the Middle East. The region's rapid economic growth and increasing trade volumes…
Cross-Border Trade and Customs Hurdles: Streamlining Logistics for Seamless Regional Commerce
Streamlining cross-border trade and overcoming customs hurdles are essential for promoting seamless regional commerce in the Middle East. The region's diverse geopolitical landscape and varying…
Navigating Geopolitical Complexity: Middle East Logistics Challenges Amidst Regional Dynamics
The Middle East faces unique logistics challenges amid the complexities of regional geopolitics and ongoing conflicts. Navigating these challenges requires careful planning, strategic partnerships, and…
Cybersecurity in Middle East Logistics: Protecting Critical Data and Digital Supply Chains
Cybersecurity is a critical aspect of Middle East logistics, as the increasing digitization of supply chains and data exchange creates new vulnerabilities and risks. Protecting…
E-Commerce Boom and Last-Mile Delivery Challenges: Adapting Logistics for Changing Consumer Behavior
The e-commerce boom in the Middle East has transformed consumer behavior, leading to a significant increase in online shopping and creating new challenges for last-mile…
Technologies transforming the future of logistics
When a commodity is ordered online or is being parceled from one place to another, generally a folk think about two persons, one who received…
Supply chain management: Asset for a product based business
Present era is completely digitized and in this digital fast moving world supply chain is acting as the best possible way of communication for all…
MANUFACTURING
Arab Aerospace Manufacturing: Taking Flight with High-Tech Innovation
Arab aerospace manufacturing is taking flight with high-tech innovation, positioning the region as a significant player in the global aerospace industry. With a focus on…
Green Manufacturing Initiatives in Arab Countries: Sustainability Meets Industry
Green manufacturing initiatives in Arab countries are gaining momentum as sustainability meets industry. With growing awareness of environmental challenges and the need to transition towards…
Localization and Resilience: Arab Manufacturing’s Pursuit of Supply Chain Innovation
Localization and resilience are becoming key priorities in the Arab manufacturing sector as companies seek to enhance their supply chain innovation. The global disruptions caused…
Smart Factories in the Arab World: Digitization Reshaping Manufacturing Landscape
Smart factories are reshaping the manufacturing landscape in the Arab world, with digitization playing a pivotal role in transforming traditional manufacturing processes. As the region…
Arab Manufacturing Renaissance: Tech-Driven Innovation Transforming Industries
The Arab manufacturing renaissance is fueled by tech-driven innovation, transforming industries across the region. As countries in the Middle East embrace the opportunities presented by…
Digital Twin Technology: Enhancing Product Development and Quality Control in the Middle East
Digital twin technology is making a significant impact on product development and quality control in the Middle East. This cutting-edge technology creates virtual replicas of…
Robotics and Cobots in Middle East Factories: The Future of Human-Machine Collaboration
Robotics and collaborative robots, known as cobots, are playing an increasingly significant role in Middle East factories, shaping the future of human-machine collaboration in manufacturing.…
Sustainable Manufacturing in the Middle East: Balancing Technology and Environmental Responsibility
Sustainable manufacturing in the Middle East is a crucial endeavor that seeks to balance technological advancements and environmental responsibility. As the region strives to embrace…
Industry 4.0 Revolution in the Middle East: Transforming Manufacturing through Technology
The Industry 4.0 revolution is transforming manufacturing in the Middle East by leveraging advanced technologies to create smart, efficient, and interconnected manufacturing processes. The region…
Additive Manufacturing in the Middle East: 3D Printing’s Impact on Innovation
Additive manufacturing, commonly known as 3D printing, has been making a significant impact on innovation and manufacturing in the Middle East. This advanced technology allows…
IOT & Manufacturing Industry
Much is being stated about Industry 4.0 and how this has changed the manufacturing landscape. Manufacturers have been able to improve operational visibility, save costs,…
Latest technologies that are going to dominate the manufacturing industry in 2021
Industry 4.0 is a new word that is emerging in the manufacturing industry and has started dominating the industry. 4.0 is a new revolution that…
Latest technologies included in manufacturing
Right from the beginning of the era, manufacturing industry is recognized for using the latest technologies. Most of the technologies are aiding manufacturing companies to…
How technology can aid manufacturing industries?
Nearly every business is evolving as a result of technological advancements. Manufacturers are continually looking for the newest and best technologies, techniques, and systems to…
Role of AR and VR in manufacturing
Manufacturing trends in 2021 will be similar, albeit “convenience” might not have been the perfect word. Manufacturing's ultimate products will remain mostly unchanged, but the…
The technologies to adopt in manufacturing industries to be in the race.
The obstacles all faced in 2020 certainly paved the way for significant progress in the coming year. As one considers the industry trends that will…
MEDICAL TECH
Telemedicine in the Middle East Revolutionizing Healthcare Delivery
Telemedicine is indeed revolutionizing healthcare delivery in the Middle East, transforming the way healthcare services are accessed and delivered in the region. Telemedicine leverages digital…
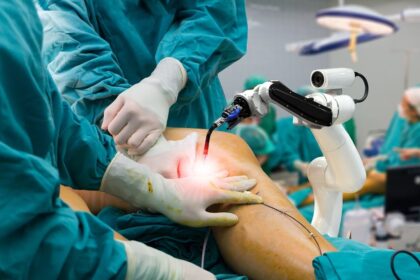
Robotic Surgery Advancements in the Middle East Precision and Innovation
Robotic surgery advancements in the Middle East have witnessed significant growth, with hospitals and healthcare institutions embracing precision and innovation in the field of minimally…
Middle Eastern Hospitals of the Future Integrating Technology for Improved Patient Care
The hospitals of the future in the Middle East are set to undergo a significant transformation, integrating cutting-edge technology to enhance patient care, improve efficiency,…
Healthcare Wearables and IoT Tracking Wellness in the Middle East
Healthcare wearables and IoT devices are gaining popularity in the Middle East as they offer new opportunities to track and monitor wellness, enabling individuals to…
E-Pharmacies and Prescription Apps The Digital Transformation of Middle Eastern Pharmacies
E-pharmacies and prescription apps are revolutionizing the pharmaceutical sector in the Middle East, ushering in a digital transformation of traditional pharmacies. These digital platforms leverage…
Culturally Sensitive Health Tech Adapting Medical Innovations to Middle Eastern Traditions
Adapting medical innovations to Middle Eastern traditions requires a culturally sensitive approach that respects the region's diverse cultural, religious, and social norms. Health tech developers…
Assistive Technologies for Special Needs Innovations in Middle Eastern Accessibility
Assistive technologies for special needs are playing a significant role in promoting accessibility and inclusivity in the Middle East. These technologies leverage advancements in AI,…
AI-Powered Diagnostics Enhancing Medical Accuracy in the Middle East
AI-powered diagnostics is revolutionizing healthcare in the Middle East by enhancing medical accuracy and efficiency. AI algorithms, when integrated with medical imaging and diagnostic systems,…
Virtual Reality Therapy in Middle Eastern Mental Healthcare
Virtual Reality (VR) therapy is emerging as a promising tool in Middle Eastern mental healthcare, offering innovative and immersive interventions for various mental health conditions.…
Bioinformatics in Genomic Medicine Decoding Middle Eastern Genetic Diversity
Bioinformatics in genomic medicine is playing a crucial role in decoding the genetic diversity in the Middle East. Genomic medicine leverages advanced computational and analytical…
Cyber-attacks on supply chain management during vaccine delivery
Cyber-attacks with a specific target- There is more to this issue than meets the eye. When considering the complete supply chain, one must consider transportation…
Medical sector to witness a great change in the upcoming decade
A study conducted over the medical sector depicts that a dynamic transformation is about to witness. Healthcare technologies are going to revolutionize the medical sector,…
RETAIL AND E-COMMERCE
Logistics Tech Disruptions: E-Commerce Delivery Challenges and Innovations in the Middle East
Logistics technology disruptions are reshaping the e-commerce delivery landscape in the Middle East. As the e-commerce market grows, meeting delivery challenges and adopting innovative solutions…
Personalization and AI in Middle East E-Commerce: Navigating Data Privacy and Enhancing Customer Engagement
Personalization and AI play a significant role in enhancing customer engagement and driving growth in Middle East e-commerce. However, navigating data privacy concerns is crucial…
Middle East Retail Tech Transformation: Overcoming Digital Adoption Challenges for Traditional Businesses
The retail industry in the Middle East is undergoing a tech transformation, driven by the growing demand for digital shopping experiences. However, traditional retail businesses…
Sustainable E-Commerce Practices in the Middle East: Balancing Growth with Environmental Responsibility
Sustainable e-commerce practices in the Middle East are crucial for balancing the region's rapid e-commerce growth with environmental responsibility. As the demand for online shopping…
Omnichannel Retailing in the Middle East: Bridging the Gap Between Online and Offline Experiences
Omnichannel retailing is becoming increasingly important in the Middle East as retailers seek to bridge the gap between online and offline shopping experiences. Omnichannel retailing…
Digital Payments and Consumer Trust: Overcoming Security Concerns in Middle East E-Commerce
Digital payments play a crucial role in driving the growth of e-commerce in the Middle East. However, consumer trust in online transactions can be hindered…
Cross-Border Commerce in the Middle East: Navigating Regulatory and Payment Hurdles
Cross-border commerce in the Middle East offers immense growth potential, but it also presents specific challenges related to regulatory compliance and payment complexities. Navigating these…
Middle East E-Commerce Boom: Addressing the Logistics Challenges of Online Retail
The e-commerce boom in the Middle East has brought numerous opportunities for online retailers and logistics providers, but it also presents specific challenges that need…
Mobile Commerce Surge in the Middle East: Meeting the Tech Demands of a Smartphone-Driven Market
The surge in mobile commerce (m-commerce) in the Middle East presents significant opportunities for businesses to tap into the region's growing smartphone-driven market. As mobile…
Middle East Payment Tech Evolution: Overcoming Digital Payment Adoption Barriers
The Middle East is witnessing a significant evolution in payment technology, with digital payment options becoming increasingly prevalent. However, there are still some barriers to…
Revolution Of Retail Industry
Retail is constantly transforming its face and now with latest four revolutions a great enhancement has been noticed in this industry. With the help of…
Trends in Retail technology
E-commerce has seen positive growth from the beginning, due to unparalleled events happening in 2020 the growth has soared higher since a year, as people…
Locus Robotics raises $150M
The E-commerce sector is growing at a fast pace and the workforce working every moment for this success is the backbone of the retail industry.…
Just ad or a story: What is more important?
This highly competitive world totally depends on show off. Audience always wishes to watch something better something creative, and day by day ad’s creativity is…
Upcoming Transformation In The Retail Sector
In the present era of technological advancements each sector is growing at a fast pace. Retail is one of those sectors that are transforming its…
How retail sector can hold their customers for the long term & reduce their expenses?
Since 2020, the world has undergone a lot of changes and a leap in the e-commerce sector. Where most of the business sectors have seen…
SMART CITY
The Future of Vertical Farming: Middle Eastern Efforts in Urban Agriculture
The future of vertical farming in the Middle East holds significant promise for addressing food security and sustainable agriculture challenges in the region. Vertical farming…
The Evolution of Smart Cities: Sustainable Urban Planning in the Middle East
The evolution of smart cities in the Middle East has been remarkable, with a focus on sustainable urban planning to address the challenges of rapid…
Tech-Enhanced Pilgrimage: Middle Eastern Cities Integrating Smart Pilgrim Services
Tech-enhanced pilgrimage is transforming the experience of religious travelers visiting Middle Eastern cities for pilgrimage purposes. Smart pilgrim services leverage technology and innovative solutions to…
Tech-Driven Water Desalination: Addressing Water Scarcity in the Middle East
Tech-driven water desalination is a critical solution for addressing water scarcity in the Middle East, where freshwater resources are limited, and the demand for water…
Smart Water Management: IoT Solutions for Conservation in Arid Middle Eastern Regions
Smart water management through IoT solutions is crucial for conserving water in arid Middle Eastern regions, where water scarcity is a significant challenge. IoT technology…
Smart Tourism: Tech-Driven Travel Experiences in Middle Eastern Destinations
Smart tourism, also known as tech-driven or digital tourism, is revolutionizing the travel industry in the Middle East, enhancing the overall travel experience for both…
Smart Manufacturing: Industry 4.0 and Robotics in Middle Eastern Factories
Smart manufacturing, also known as Industry 4.0, is transforming the industrial landscape in the Middle East, with an increasing focus on automation, robotics, and advanced…
Smart Home Solutions: Integrating IoT for Comfort and Efficiency in Middle Eastern Homes
Smart home solutions are becoming increasingly popular in the Middle East, as they offer convenience, comfort, and energy efficiency to homeowners. These solutions leverage the…
Smart Energy Grids: Integrating Renewable Energy Sources in Middle Eastern Power Systems
Integrating renewable energy sources into smart energy grids is a promising approach to enhance the sustainability and efficiency of power systems in the Middle East.…
AI-Driven Conservation Drones: Monitoring Wildlife Habitats in the Middle East
AI-driven conservation drones can play a vital role in monitoring and protecting wildlife habitats in the Middle East. These drones leverage artificial intelligence (AI) and…
The first pillar of a smart city – Institutional development
The concept of the smart city differs from person to person. For some, a city flourishes with its institutional growth, and for some, an excellent…
The second pillar of a smart city – Development of physical infrastructure
When all focus is on building your smart city, how can we negate the role of the development of physical infrastructure? The purpose of physical…
The fourth pillar of a smart city – Development of economic infrastructure
To develop an ideal city that will the future filled with opportunities, it requires a basic economic infrastructure. There are two fundamental goals that any…
Smart City and its growing impact
The idea of a smart city has become extremely important when it comes to the context of defining the strength, performance, facility, demography and urban…
The third pillar of a smart city – Development of social infrastructure
Imagine a city has everything that it takes to be called as a smart city; the basic amenities, physical infrastructure, well-planned governance institution and all.…
Moscow introduces blockchain e-voting to overhaul smart-city plans
The Moscow local government is looking to implement an often-discussed idea: an e-voting system built on blockchain. While many issues have been raised (particularly in…
TECHNOLOGY
Cultural Heritage Preservation: Government Efforts to Safeguard History and Identity in the Middle East
Cultural heritage preservation is of paramount importance in the Middle East, given the region's rich history and diverse cultural identity. Middle Eastern governments recognize the…
IoT and Energy Efficiency Middle Eastern Innovations for a Sustainable Future
IoT (Internet of Things) is playing a significant role in driving energy efficiency and promoting sustainability in the Middle East. As the region faces growing…
Blockchain in Insurance: Middle East’s Journey Toward Secure and Transparent Transactions
Blockchain technology is transforming the insurance industry in the Middle East, offering secure and transparent transactions that improve trust, efficiency, and customer experience. Blockchain's decentralized…
AI and Financial Inclusion: Bridging the Banking Gap in the Middle East
AI is playing a transformative role in advancing financial inclusion and bridging the banking gap in the Middle East. With the region's diverse population and…
Cybersecurity in Middle East Logistics: Protecting Critical Data and Digital Supply Chains
Cybersecurity is a critical aspect of Middle East logistics, as the increasing digitization of supply chains and data exchange creates new vulnerabilities and risks. Protecting…
Innovations In The Automotive Industry
Information-oriented technologies play a key role shortly in the automobile industry, according to a comprehensive analysis of industry trends to expect forward to in 2021.…
Rethinking healthcare services with AI-enabled physicians
While many physicians have been cautious about artificial intelligence in clinical settings in the past, in today's fast-changing healthcare sector, many are considering how AI…
Strategies to Improve Public Sector Labour Productivity
Labour productive is an essential element for any organization to perform well and achieve intended goal. To keep your organizational working procedure effective and accurate,…
A complete guide of BFSI
BFSI stands for Banking, Financial and Insurance sector. Generally, it represents a major portion of the Indian economy which comprises of all Banking Insurance and…
Empower your business with the next level of Cloud Computing
From Applications to storage and processing power, under the umbrella of cloud computing, it includes the process of delivery of on-demand computing services. Cloud computing…
Protect your Digital Infrastructure from cyber attacks
Cyber Security is a term used to define every technology, process, and practice designed to protect networks, devices, and data from any kind of cyberattacks.…
Topic of interest in blockchain technology
Bockchain technology is gaining popularity in recent years and is able to attain attention of techies. It can be defined as a digital ledger having…